Author: Denis Avetisyan
A new framework harnesses the power of large language models to automate the process of scientific model discovery, moving beyond traditional methods.
![ModelSMC automatically discovers models from textual problem formulations and context data by iteratively refining an initial model-inspired by Sequential Monte Carlo methods that approximate distributions via weighted particles-to sample high-density regions of the model posterior [latex] p(m|\bm{x}_o) [/latex], effectively approaching the underlying data-generating process through model propagation via large language model sampling and weighting based on likelihood evaluation.](https://arxiv.org/html/2602.18266v1/x1.png)
The paper introduces ModelSMC, a probabilistic programming approach using Sequential Monte Carlo and Bayesian inference to enable robust and interpretable scientific discovery with LLMs.
Automated scientific model discovery currently relies on heuristic procedures that lack a unifying theoretical foundation. This limitation motivates the work presented in ‘A Probabilistic Framework for LLM-Based Model Discovery’, which recasts the problem as probabilistic inference, enabling a more robust and interpretable approach. Specifically, we introduce ModelSMC, an algorithm leveraging Large Language Models and Sequential Monte Carlo to represent and refine candidate mechanistic models as particles weighted by their explanatory power. Could this probabilistic lens unlock a new generation of LLM-driven tools for accelerating scientific progress and uncovering hidden mechanisms within complex systems?
The Challenge of Mechanistic Modeling
The creation of mechanistic models, those that attempt to explain how a system functions rather than merely predicting what it will do, has historically been a deeply human endeavor. It demands not only a robust understanding of the underlying principles governing the system – be it a biochemical pathway, a climate pattern, or an economic market – but also significant manual effort in formulating hypotheses, designing experiments to test them, and then painstakingly translating the results into a mathematical framework. This process frequently necessitates years of dedicated research by specialists with extensive domain expertise, a substantial investment of time and resources, and carries an inherent risk of cognitive biases influencing model construction. Consequently, the pace of discovery can be limited, and the potential for overlooking alternative explanations remains a persistent challenge within scientific modeling.
The construction of mechanistic models, while crucial for understanding complex systems, frequently encounters significant bottlenecks that impede the pace of discovery. Traditional methods demand extensive expertise, requiring researchers to painstakingly define relationships and parameters – a process that can be both time-consuming and financially burdensome. Moreover, this manual approach introduces the potential for unconscious bias, as researchers’ preconceptions can inadvertently shape the model’s structure and interpretation. Consequently, the development of accurate and reliable models can be dramatically slowed, limiting the ability to quickly test hypotheses, explore new phenomena, and ultimately, accelerate scientific progress across diverse fields.
Automated Inference via Sequential Monte Carlo
Sequential Monte Carlo (SMC) is a class of particle-based methods used for Bayesian inference and model selection, particularly effective when dealing with non-linear and non-Gaussian systems. The core principle of SMC involves representing probability distributions with a set of weighted particles, each representing a possible state of the system. These particles are propagated through time via a prediction step, followed by a weighting step based on observed data. The weights, proportional to the likelihood of the data given the particle’s state, are then normalized, and a resampling step eliminates particles with low weights, concentrating computational effort on more promising regions of the state space. This iterative process allows SMC to approximate posterior distributions and estimate model evidence, enabling probabilistic model comparison and uncertainty quantification. [latex] p(\theta|x) \approx \sum_{i=1}^{N} w_i^{(t)} \delta(\theta – \theta_i^{(t)}) [/latex] where θ represents the model parameters, [latex] x [/latex] the observed data, and [latex] w_i^{(t)} [/latex] the normalized weights of the particles at time step [latex] t [/latex].
Sequential Monte Carlo (SMC) utilizes a particle-based approach to approximate probability distributions that are analytically intractable. This method represents the distribution as a set of weighted samples, termed “particles,” each representing a possible state of the system. These particles are propagated through time or model space via a resampling and weighting scheme. Particles with higher weights, indicating greater probability, are duplicated, while those with lower weights are eliminated, effectively focusing computational effort on the most promising regions of the parameter space. This process allows SMC to effectively explore complex, high-dimensional distributions and identify model candidates with high posterior probability, even in scenarios where direct evaluation of the probability density function is impossible.
Integrating Sequential Monte Carlo (SMC) with Large Language Models (LLMs) enables automated model discovery by utilizing the LLM’s code generation capabilities to create candidate model implementations. SMC provides the probabilistic framework for evaluating these LLM-generated models; the LLM synthesizes a model, SMC assesses its performance against observed data via weighted resampling of a particle set, and this feedback loop iteratively refines the generated models. The LLM can then generate variations on promising models, effectively exploring the model space and identifying high-likelihood candidates without explicit human intervention. This process allows for the automation of tasks previously requiring substantial expert knowledge in both modeling and probabilistic inference, facilitating the discovery of novel models tailored to specific datasets and objectives.

Quantifying Model Fit and Efficiency
Model evaluation relies on quantifying how well a model aligns with observed data, and this alignment is typically assessed using metrics such as ResamplingWeight. ResamplingWeight effectively represents the probability of a model being selected during a Sequential Monte Carlo (SMC) process, given the observed data; higher weights indicate better explanatory power. This metric is derived from the likelihood of the observed data given the model’s parameters, and is used to differentiate between models that adequately describe the data and those that do not. The accumulation of ResamplingWeight across multiple iterations of SMC provides a robust measure of a model’s overall fit, allowing for comparative analysis of different model structures and parameterizations.
Neural Likelihood Estimation (NLE) and Neural Posterior Estimation (NPE) provide computationally efficient alternatives to traditional methods for approximating likelihood and posterior distributions, which are essential components of Sequential Monte Carlo (SMC) algorithms. Traditional likelihood calculations can be intractable for complex models, requiring significant computational resources. NLE utilizes neural networks to directly learn the likelihood function from data, bypassing the need for explicit analytical calculations. Similarly, NPE employs neural networks to approximate the posterior distribution, enabling efficient sampling. By leveraging the representational power of neural networks, NLE and NPE significantly reduce the computational cost associated with SMC, facilitating model inference and parameter estimation in scenarios where exact calculations are impractical.
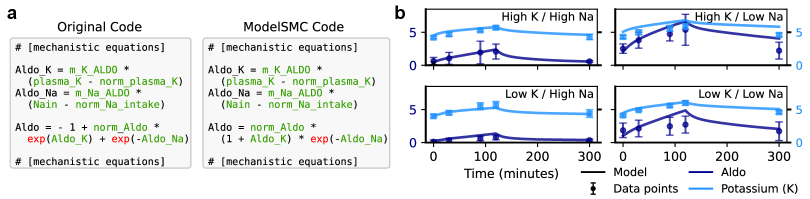
ModelSMC’s efficacy in identifying plausible models has been validated across three distinct biological systems. The HodgkinHuxleyModel, a well-established computational model of neuron electrical properties, was successfully analyzed, alongside the SIRModel, which simulates the spread of infectious diseases within a population. Furthermore, the PharmacologicalKidneyModel, detailing the complex functional mechanisms of the kidney, also demonstrated successful model identification using ModelSMC. This cross-system validation confirms the broad applicability of the method beyond any single specific biological context, indicating robustness in its ability to infer meaningful parameters from diverse datasets.
The computational efficiency of the automated model selection process is quantified using TokenUsage, which measures the number of tokens processed during model evaluation. Across the SIR, Hodgkin-Huxley, and PharmacologicalKidneyModel systems, ModelSMC demonstrates performance comparable to baseline methods like FunSearch+ and ModelSMC NN=1, achieving similar results within a comparable token budget. This indicates that ModelSMC achieves a balance between model accuracy – as measured by Negative Log Likelihood – and computational cost, making it a viable option for automated scientific modeling.
ModelSMC demonstrates performance competitive with existing methods, as evidenced by achieving comparable Negative Log Likelihood (NLL) scores across the Hodgkin-Huxley, SIR, and PharmacologicalKidney models. This competitive performance is maintained within a similar token usage budget as comparison methods, including FunSearch+ and a single-layer Neural Network instantiation of ModelSMC (ModelSMC NN=1). The consistency of NLL scores, given comparable computational cost measured by token usage, indicates that ModelSMC provides an efficient and statistically viable alternative for model selection and inference tasks across these diverse systems.
![Across three dynamical systems-a SIR epidemiological model, the Hodgkin-Huxley model, and a pharmacological kidney model-ModelSMC and FunSearch+ consistently outperform Model SMCNN=1, as measured by negative average log-likelihood using both marginalized resampling weights ([latex]Eq.5[/latex]) and parameter estimates ([latex]Eq.E-{40}[/latex]).](https://arxiv.org/html/2602.18266v1/x5.png)
The Dawn of Automated Scientific Discovery
The automation of model discovery represents a paradigm shift in scientific investigation, enabling researchers to systematically evaluate a significantly broader spectrum of potential hypotheses than previously feasible. Traditionally, the formulation and testing of models relied heavily on human intuition and expertise, inherently limiting the exploration of model space. By employing algorithms to navigate and assess various model configurations, this automated approach circumvents these limitations, potentially uncovering previously overlooked relationships and accelerating the pace of discovery. This is not merely about faster computation; it’s about expanding the search itself, moving beyond pre-conceived notions to identify models that accurately reflect underlying phenomena and ultimately drive scientific progress across diverse disciplines.
This innovative framework transcends the limitations of domain-specific analysis by offering a versatile platform applicable to diverse scientific fields. From astrophysics and materials science to drug discovery and climate modeling, the system’s core principles – automated hypothesis generation and sequential model refinement – remain consistent, yet readily adapt to the unique datasets and governing equations of each discipline. This adaptability stems from its reliance on general computational techniques rather than specialized algorithms, allowing researchers to leverage the same infrastructure for analyzing vastly different phenomena. Consequently, it facilitates knowledge generation not just within a single area of study, but across the entire scientific landscape, potentially uncovering unexpected connections and accelerating breakthroughs in multiple domains.
Recent advancements showcase a compelling synergy between large language models and Sequential Monte Carlo methods, exemplified by the FunSearch framework, to unlock novel avenues for scientific discovery. This approach transcends traditional model-building by enabling a form of ‘creative’ exploration of the model space – rather than testing pre-defined hypotheses, the system actively proposes and evaluates models based on the data. By leveraging the generative capabilities of LLMs to suggest model structures and the robust sampling of SMC to refine parameters, researchers can navigate complex landscapes and identify solutions previously overlooked. This is not simply automation; it’s a paradigm shift towards allowing algorithms to contribute to the formulation of scientific theories, potentially accelerating breakthroughs in fields ranging from physics to biology by discovering unexpected relationships within data.
The integration of large language models with Sequential Monte Carlo methods significantly refines the process of parameter estimation, yielding models of demonstrably higher accuracy and reliability. Traditional methods often struggle with complex, high-dimensional datasets, leading to imprecise or unstable results; however, this framework leverages the LLM’s capacity to intelligently guide the SMC algorithm towards promising parameter configurations. This directed search reduces computational cost and minimizes the risk of converging on local optima, allowing for a more thorough exploration of the model space. Consequently, researchers can obtain parameter sets that not only fit the observed data effectively, but also generalize well to unseen data, bolstering the predictive power and robustness of scientific models across diverse fields.
The pursuit of robust model discovery, as outlined in this work, demands a commitment to demonstrable correctness. Any ambiguity introduces the potential for abstraction leaks, undermining the integrity of the derived scientific understanding. This echoes Linus Torvalds’ sentiment: “Most programmers think that if their code ‘works’ then it’s good enough. I disagree.” The ModelSMC framework, by explicitly framing discovery as probabilistic inference, seeks to move beyond merely ‘working’ solutions, striving instead for provable, mathematically sound models. The probabilistic approach isn’t simply about finding a model, but quantifying the confidence in its correctness, aligning with a philosophy that prioritizes verifiable truth over pragmatic functionality.
What’s Next?
The presented framework, while a step towards automating scientific discovery, ultimately shifts the burden of proof. It does not create understanding; it merely accelerates the process of hypothesis generation and evaluation. Let N approach infinity – what remains invariant? The fundamental need for rigorous validation, grounded in first principles, does not diminish with increased computational power. The elegance of a model is not determined by its ability to fit data, but by its internal consistency and predictive power beyond the training set – qualities that probabilistic inference alone cannot guarantee.
A critical limitation lies in the LLM itself. These models, trained on correlation, not causation, introduce a subtle, yet pervasive, bias. The framework’s reliance on LLM-generated proposals necessitates careful consideration of the prior distribution imposed by the language model – a prior reflecting the biases inherent in the corpus from which it learned. Future work must address this, perhaps through the development of LLMs specifically trained on scientific literature, and designed to explicitly represent uncertainty.
The true challenge, however, remains not in automating the process of discovery, but in formalizing the criteria for scientific merit. A system can efficiently explore a vast hypothesis space, but it cannot, of itself, determine which hypotheses are truly meaningful, or which represent genuine advances in understanding. The pursuit of automated science, therefore, requires not only sophisticated algorithms, but also a deeper philosophical inquiry into the nature of scientific knowledge itself.
Original article: https://arxiv.org/pdf/2602.18266.pdf
Contact the author: https://www.linkedin.com/in/avetisyan/
See also:
- ‘Project Hail Mary’s Unexpected Post-Credits Scene Is Worth Sticking Around
- Total Football free codes and how to redeem them (March 2026)
- Limbus Company 2026 Roadmap Revealed
- The Division Resurgence Specializations Guide: Best Specialization for Beginners
- After THAT A Woman of Substance cliffhanger, here’s what will happen in a second season
- Brawl Stars Sands of Time Brawl Pass brings Sandstalker Lily and Sultan Cordelius sets, along with chromas and more
- Brawl Stars Brawl Cup Pro Pass arrives with the Dragon Crow skin and Chroma, unique cosmetics, and more rewards
- Clash of Clans April 2026 Gold Pass Season introduces a Archer Queen skin
- XO, Kitty season 3 soundtrack: The songs you may recognise from the Netflix show
- Gold Rate Forecast
2026-02-23 18:14